Astonishing biological patterns surround us from the stripes on a zebra to the hair follicles on our skin. In fact, you might devote a significant part of your life getting rid of body hair, but you would be amazed by the way that nature puts them there in the first place. You could even call it a work of art. It comes as no surprise that pattern formation has been an interesting corner of biology, especially in the developmental biology field because it is in the embryo where the patterns of our future physiology are laid down. In fact, much of developmental biology is concerned with pattern formation by which initially equivalent cells generate different functions and morphologies. Trying to understand the principles behind pattern formation guides us to understand how organisms form with tissues in the right place at the right time.
Pattern formation has been studied for years, famously with Alan Turing having an influential impact on the field in 1952 (1). He devised a mathematical framework called the reaction-diffusion Turing networks which could explain how patterns arise from initially uniform conditions. Using various mathematical rules and relationships, it is possible to simulate the formation of many familiar patterns such as stripes and dots. However, it has been difficult to determine whether Turing networks actually drive biological pattern formation. For instance, these mathematical models are simplified – simplification and assumptions are of course needed in a model to compute the outcome and to understand the core principles of a system. They are useful for making insightful and sometimes counterinitiative predictions that can then be experimentally tested. So, there must be a two-way conversation between the model and the biology to determine whether the mathematical rules can explain a biological system.
Turing fully acknowledged the limitations of his model; ‘This model will be a simplification and an idealisation, and consequently a falsification’ (1). Furthermore, statistician, George E. P. Box, said that ‘all models are wrong but some are useful’, which is a succinct way of saying that models cannot fully explain a system but they can provide insights regarding a specific question you are asking. Indeed, the Turing model has been extremely influential as a conceptual framework. In some cases, it has given biologists a starting point to probe their questions concerning pattern formation.
So how do the reaction-diffusion networks function? Suppose you have two interacting diffusible molecules and one is an activator and one is an inhibitor. Here, the activator enhances the expression of itself as well as introducing the expression of an inhibitor. These molecules have different diffusion rates to generate different concentrations of the inhibitor and activator across a cellular field. Specifically, the inhibitor diffuses faster than the activator and therefore, as soon as the inhibitor is produced it diffuses away. This generates an array of activation peaks where the concentration of the inhibitor is not enough to inhibit the activator (figure 1). The idea is that these diffusible molecules, known as morphogens, change the behavior of cells in order to generate morphological patterns. This is a very simple case, and the Turing networks can manifest in alternative ways depending on the context. For instance, in biological examples, they often constitute a three component Turing network. This has been shown during digit patterning in the limb and during the patterning of various epidermal appendages (e.g. hair follicles in mammals and skin denticles in sharks) (2) (3) (4).
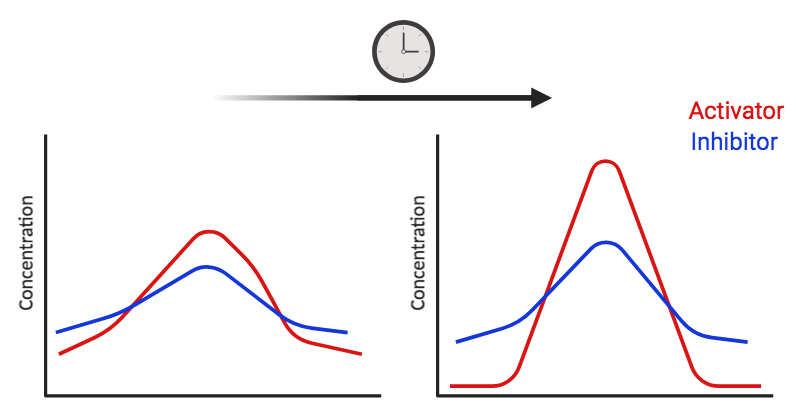
Figure 1. Simple depiction of Turing reaction-diffusion networks with two interacting diffusible molecules.
As mentioned above, one particular example where a reaction-diffusion mechanism can partly explain pattern formation is during hair follicle development. A reaction-diffusion interaction network between three molecules, bone morphogenetic protein–fibroblast growth factor–wingless-related integration site (BMP–FGF–WNT), has been shown to be sufficient to generate a periodic epidermal (outer skin layer) pre-pattern (3). Here, the pre-pattern refers to the differential distribution of diffusible molecules that will later generate a morphological structure. However, the reaction-diffusion model is not the only model that could generate the pattern of hair follicles. The same results can also be generated from a mesenchymal self-aggregation model (figure 2). In this model, the cells themselves are the agents of pattern rather than diffusible molecules. Cells would aggregate to form the precursors of a hair follicle but at the same time there would be a lower density of cells surrounding the aggregate. In fact, it was found that the dermal cells underneath the epidermis can self-organise by such a mechanism given the right conditions.
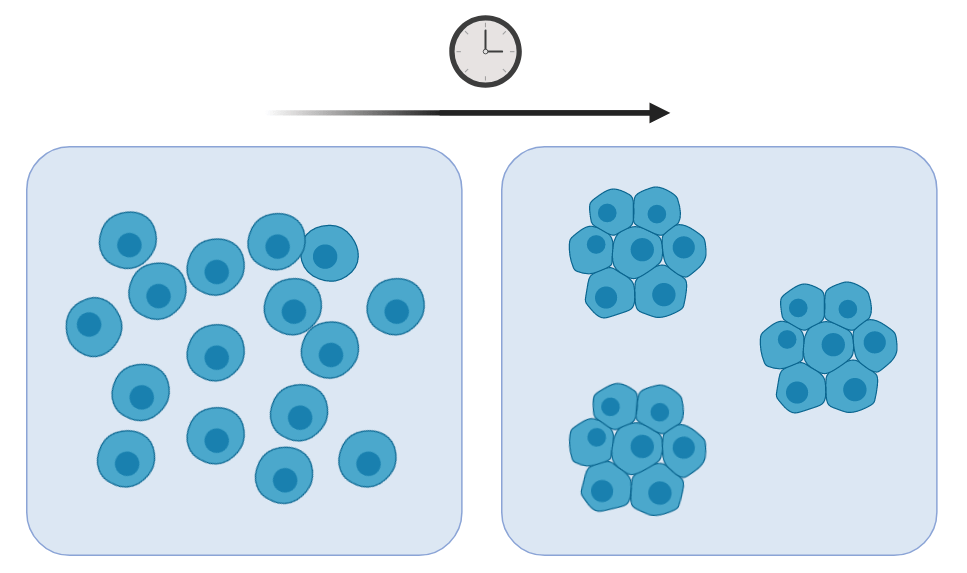
Figure 2. Mesenchymal self-aggregation model. Cells reorganise into clusters over time to generate areas of high cell density and areas of low cell density in the surrounding environment.
Putting the two models together, it was shown that a reaction-diffusion mechanism is deployed in the epidermis and this acts as a molecular pre-pattern. The dermal cells below then aggregate towards this pre-pattern to generate cellular rearrangements which ultimately determine the location of hair follicles (figure 3). Essentially, the dermis has the ability to self-organsise through cell aggregation but this organisation is initially instructed by a reaction-diffusion patterning above it. This is a fascinating example of where two patterning mechanisms intersect to generate biological form.
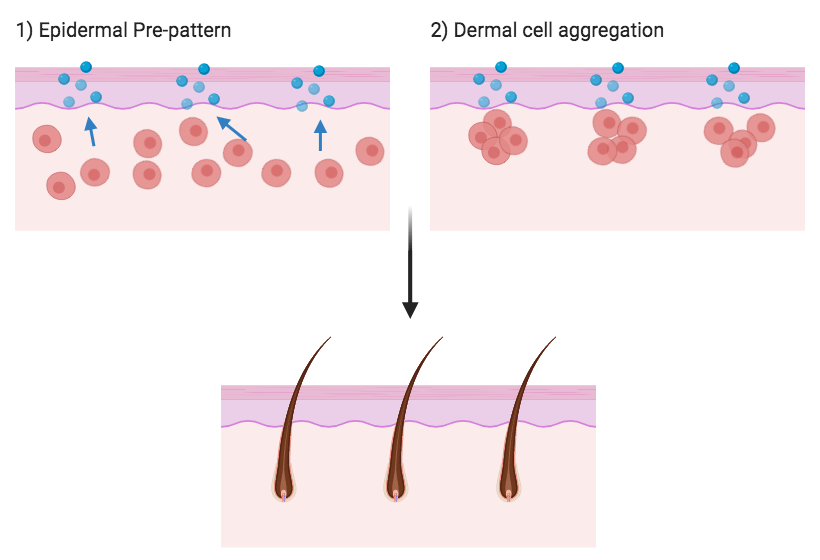
Figure 3. Integration of the reaction-diffusion patterning mechanism in the epidermis which acts as a pre-pattern to instruct the mesenchymal cell aggregation in the dermis below.
Concluding remarks
Hair follicle patterning is a wonderful example of the beauty and complexity of developmental processes; tissues must interact to coordinate the development of structures so that they form in the right place with the correct sizes. In addition, the different diffusible molecules and their varying concentrations are interpreted by cells to generate different cellular behaviors which ultimately translate into morphological structures (figure 3). Such behaviors include changes in cell migration and cell adhesiveness. In accordance with this, cells become appropriately specialised to generate different cell types later on in development. With the advancement of new imaging tools, genetic manipulation, and mathematical modelling, it is an exciting time for developmental biology. Who knows what might be hiding behind the scenes?
References
1) Turing, A. M. (1952). The chemical basis of morphogenesis. Philosophical Transactions of the Royal Society (part B), 237 (1952), pp. 37-72
2) Raspopovic, J., Marcon, L., Russo, L., Sharpe, J. (2014). Digit patterning is controlled by a Bmp-Sox9-Wnt Turing network modulated by morphogen gradients. Science 345, 566–570.
3) Glover, J.D., Wells, K.L., Matthäus, F., Painter, K.J., Ho, W., Riddell, J., Johansson, J.A., Ford, M.J., Jahoda, C.A.B., Klika, V., Mort, R.L., Headon, D.J. (2017). Hierarchical patterning modes orchestrate hair follicle morphogenesis. PLoS Biol 15, e2002117.
4) Cooper, R.L., Thiery, A.P., Fletcher, A.G., Delbarre, D.J., Rasch, L.J., Fraser, G.J. (2018). An ancient Turing-like patterning mechanism regulates skin denticle development in sharks. Sci. Adv.
Figures made courtesy of BioRender
Featured image taken by Frida Bredesen, https://unsplash.com/photos/c_cPNXlovvY
